Greetings from the Unsupervised Learning HQ!
I have a new project dropping this week that I’m excited to invite you to check out. If you’re interested in understanding your own ancestry, I think this is kind of the killer upgrade. Details follow at length below.
But first, another big thanks to all the free subscribers who have been upgrading to paid, and a couple of announcements. Also, for what it’s worth, as a minority-identifying (at least this week) content producer, I’ve been making it known that I will be happy to relieve Andrew Sullivan of some of the monetary spoils of his white privilege on Substack. With his 17,000 subscribers, the gap between us is a major issue of equity. I’ll keep you posted.
Over the past week, I released a two-part “deep dive” into the latest Indian ancestry findings and their historical and social relevance. This is a long read (~10,000 words total) and permanently paywalled, but I can sincerely recommend it!
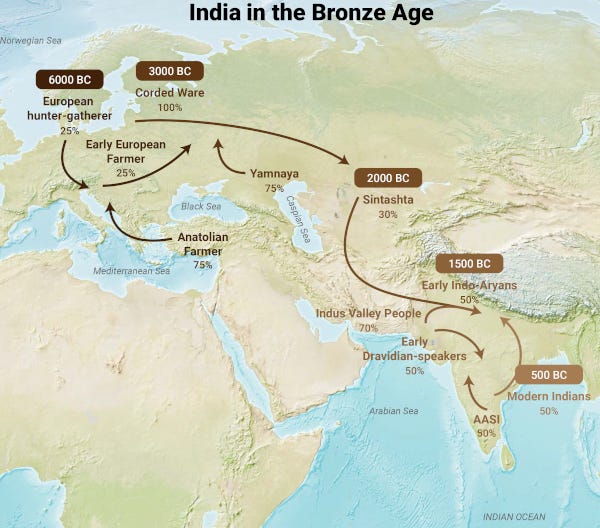
Stark Truth about Aryans: a story of India
Stark Truth about Humans: a story of India
Also, last week I had a chance to chat with Ramesh Ponnuru on the podcast about the pro-life movement in America. This week I’ll be posting my conversation with John Hawks about his life in paleoanthropology.
My latest two podcasts to come out from behind the paywall are with Armand Leroi on mutation and Aristotle and Alina Chan on the “lab leak” hypothesis. Reviews on the free content make a difference for me, so if you’re so inclined, drop me a positive review on Apple Podcast and Stitcher.
And lastly, are you following Colin Wright and doing your part to make Scott Alexander feel welcome at his new Substack perch? Equity matters, but so does interesting content.
DIY your DTC DNA: an invitation to join me live this Weds. Jan. 27
The screenshot above is from my 23andMe results. The inference that my ancestry is from "Chittagong Division" is 100% correct. More precisely, my family is from Comilla. There are some cases where consumer genomics tells you exactly what you know, and this is one of those. 23andMe has an excellent method to infer ancestry, and the power of a massive database. If you want to see how they do it, check out their new preprint. It's pretty fancy.
What would be more informative is if they let me see what it means to be from Chittagong Division compared to other Bengalis. Or to be Bengali compared to other South Asians. You probably already know your recent history through basic family genealogy, but what do these results tell you about your deep history and your relatedness to global populations?
Of course, realistically even the largest direct-to-consumer genomics companies can only deliver so much, because they are simultaneously serving millions of customers. A custom approach isn’t a feasible ask, even if it is what many consumers are longing for. Little surprise then that people have been reaching out to me about anomalies in their results for over a decade. People like me who got no value-added information (as in: they already know where their great-grandparents were born) reach out, too. We’re driven to know more.
That is why this week I'm offering my first workshop through Speakeasy titled Analyze Your 23andMe and Ancestry Data (and N.B. it didn’t make the title but I’ve prepared everything to work right on results from Family Tree DNA as well). It's Wednesday, Jan 27, 5pm PT/8 pm ET.
If you’re curious, let me walk you through how it will work. You’ll arrive in class with your (or a friend’s or relative’s) 23andMe, Ancestry or Family Tree DNA raw data. Before class, you’ll get a zip folder with all the files and utilities you need. You’ll have downloaded R & R Studio if you don’t already have them (I include instructions, but don’t worry, this is quick and easy!). And you’ll want to decide whether to Zoom into the meeting on one device and access your data on the other (or work on a two-screen setup if you’re on a desktop); neither is essential, but I’d consider them nice to have.
At that point, you’re ready for the workshop. And we’ll get straight into digging into your data. You can come to class with a question or questions about your ancestry. Or I can help you zero in on what might be interesting given what you already know.
Over the course of the hour, I’ll guide you through the use of three tools. No lecture, just hands-on doing, with your own actual data. Two of your tools are open-source utilities written by academics for their colleagues that have been in wide use in genomics for over a decade. Long-time readers of my blog will recognize Plink and Admixture whether they are into genetics or not because I’ve referenced them thousands of times over the years.
In addition to easing you right into using these core tools of the trade (without any of the usual slow initial learning curve), I have built an automated pipeline just for participants in the class. This is your third tool, which will save you untold startup hours no matter who you are. I’ve created an automated workflow so that you can input your raw data from any of those three DTC genetics companies and analyze it (including automatically generating formatted output) against your choice of reference populations (a library of which I’ve also prepared for you) and 2. automatically plot and visualize your results in a flexible, customizable format.
I have written all the scripts to create a custom, automated pipeline. This draws on my years of experience using these tools and guiding others. Then, the bespoke reference population library of human genomes I’ve curated for this purpose instantly equips you to measure relatedness to any branch or branches of humanity (you get 5000 human genotypes culled from public datasets and selected to represent 84 distinct populations, on a quarter of a million markers (SNPs).
And in class I will teach you and guide you through using your toolkit in real time. Getting started in Plink and Admixture can entail hours of trouble-shooting and false starts. A decade into using them, I know the quirks and idiosyncrasies of these programs all too well; and that’s why I’ve built a pipeline that allows you to leapfrog over those slow early steps and get right into your (or any) genetic data. Building and merging a reference panel from publicly available sources is time-consuming and a headache and the individuals don’t come clearly labeled by population. I’ve got everything ready and clearly identified for you. With these two headstarts, and lots of pointers about best practices along the way, you’ll be asking and answering (and outputting visualizations of) your own questions in your first hour.
By the end of the workshop, you'll have the skills and the tools to analyze genotypes against world population data. You’ll be able to use Plink, Admixture and the pipeline I’ve created for you on any DTC genomic results from those main three companies. You’ll have both the curated reference library and the pipeline I built for you... And the know-how to use them to your ends. You’ll also have reference cheat-sheets on how to do everything you tried in the workshop (I don’t want you to have to take notes when you can be learning by doing!)
Who is this for? You. I promise. My goal with this project is to make it accessible and easy for everyone with basic personal computing literacy. Not programming, not command line, not R. Just be comfortable on your computer. (And for this iteration, you need to be on a Mac or Ubuntu/Linux OS. I’m still working out a kink in Windows, so email me if you’d like to try it on Windows once I get that working.)
I want to reach people who aren’t geneticists. I want to reach people who assume they can’t do this. I want to show curious people who have never heard of any of the tools I’m naming that they can still delve into their DNA on their own terms the first time they sit down and try.
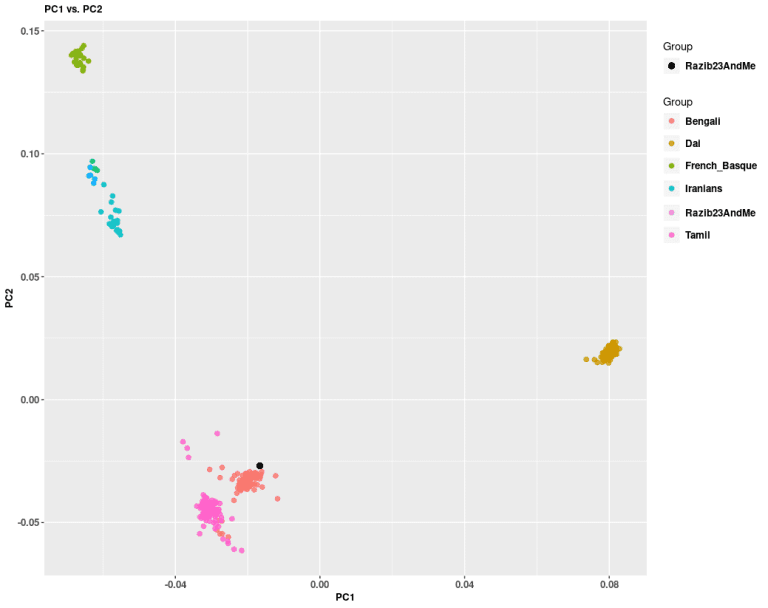
But enough about you! Let’s get back to me and what I did with my 23andMe results. To answer the question of what it means to be "Bengali from Chittagong" I analyzed myself against only a few populations and compared myself to the Bengalis in the 1000 Genomes.
The PCA immediately shows that as someone from eastern Bengal, I'm on the edge of the eastern Bengali cluster. I have a lot of East Asian ancestry. To be from Chittagong Division means to be 10-20% East Asian.
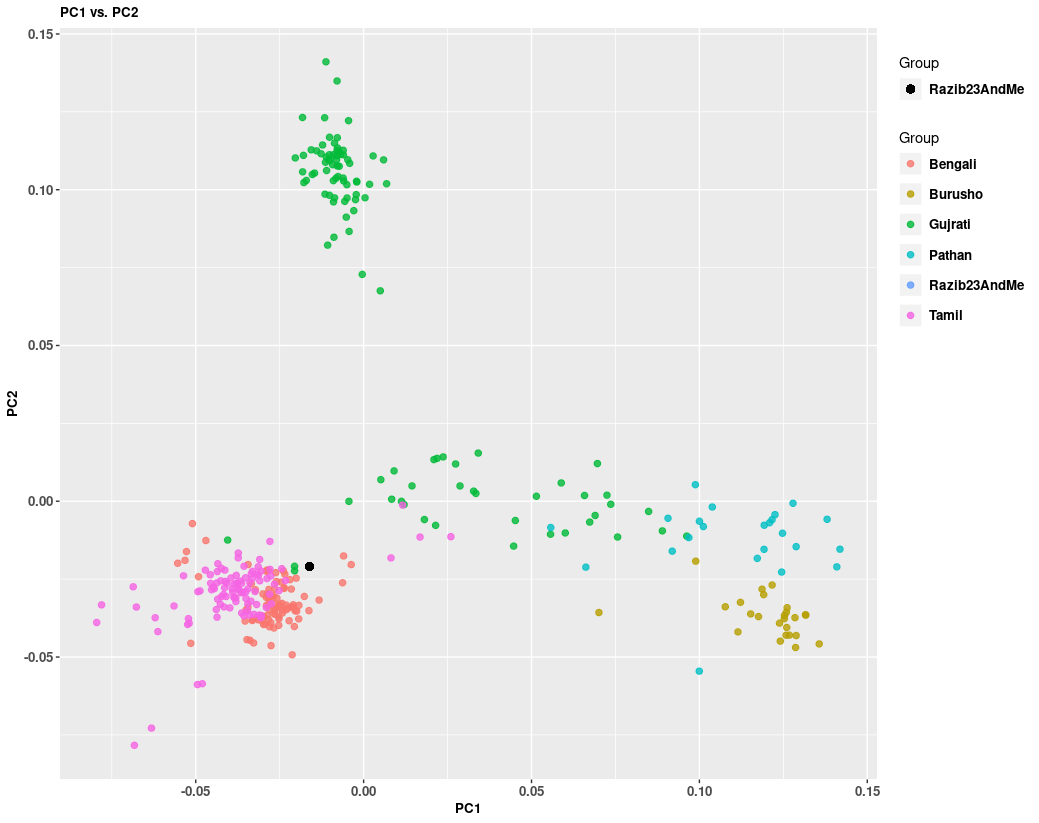
Let's rerun this with more South Asian populations for context and zoom in:
Again, you see I'm on the edge of the Bengali cluster. The same general result, but a different context. You can see here that Bengalis are somewhat different from Tamils, but closer to them than other Indian populations in the west and north of the subcontinent.
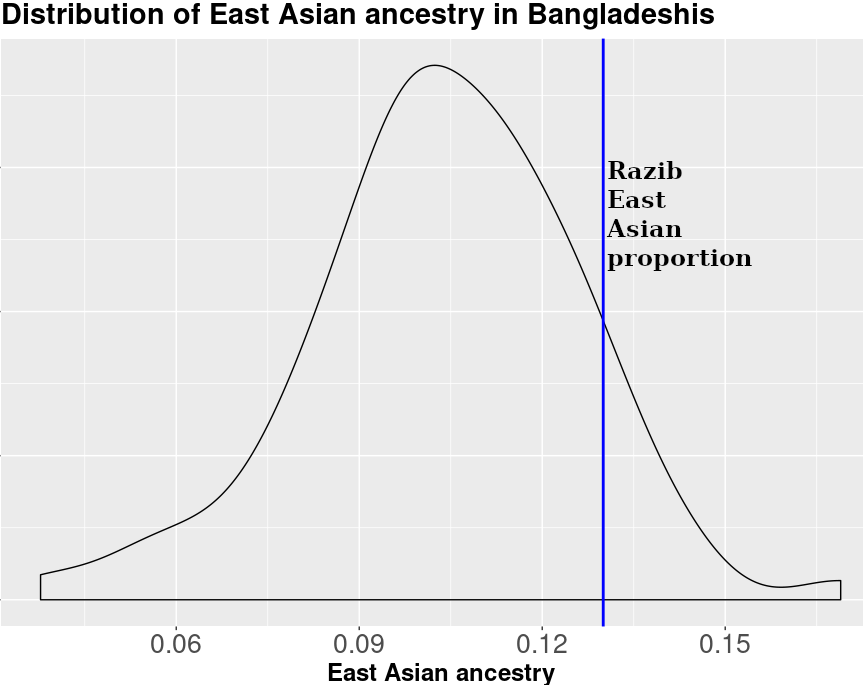
I've seen plenty of other Bangladeshis and Bengalis on 23andme. Many of us get pretty generic "You are 85-99% Bengali" interpretations. But that masks the reality of variable East-Asian ancestry across Bengal (more in the east, less in the west).
It's good to check with another method, so I used ADMIXTURE, and ran the Bangladeshis with Tamils (useful to represent South Indian for the purposes of my exploration), Dai (East Asian), and Iranians, as the reference populations. Out of 83 Bangladeshis in the sample, I'm ranked 11th in percent Iranian and seventh in percent Dai. This aligns with what we see in the PCA. I have less "Tamil" ancestry proportion. What does that mean? Basically, I have less generic Indian ancestry than the average South Asian person.
How long did it take me to figure this out? About 15 minutes. The methods of simple data exploration aren't difficult or necessarily time-consuming. They're just about the initial set-up time (which my course helps you leapfrog over) plus repetition and iteration.
This week is my first time running this, and I'm pretty excited. I trust the turnout this first time will be light (or perhaps we should say "select"). But these tools are something I’m eager to share. I want to equip people to be able to pose their own questions and help themselves, their family and friends get more out of their data. So in addition to what will probably be a nice student-teacher ratio, you’ll be helping me shape this to empower as many laypeople as possible.





Hi Razib, do you plan to run this workshop again? Or, do you have scripts/libraries posted somewhere? Thank you
When will the next one be? I missed the last one unfortunately.